
 AMINE
AMINE
About AMINE
AMINE method makes it possible to identify relevant gene modules by simultaneously taking into account the differential activity of genes and their relationship. To do this, the algorithm needs data concerning the differential activity of genes and a network of biological interactions between genes within the organism under study.
Input
P-values, representing the evidence of change of gene expression across experimental conditions, are specified in a file uploaded via the input field "Differential expression data". At a minimum, the data is composed of a tabular file consisting of two columns: one containing the name of the gene and the other, the associated p-value. A more convenient way to indicate the data to be used is to simply specify the result file produced by differential expression analysis methods such as DeSeq2 or edgeR. The input file can be in csv (comma-separated values) or xlsx format.
The two following input fields are used specify the columns containing the needed data. "Column with gene names" must indicates the column that contains the gene names while "Column with p-values" contains the index of the column with p-values. It should be noted that the columns are numbered starting at 1, which means that 1 is the index of the first column of the table.
It is possible to focus the search on modules containing specific genes. A file containing the list of these "favorite" genes (one gene name per line) can be specified with the input field "Favorite genes".
The execution of the algorithm can take several hours depending on the server load and the size of the data to be analyzed. In order not to force the user to stay logged in on the input page, we ask for an email address, which will be used to notify her of the completion of the task and provide the address where the results are available.
example of an input file :
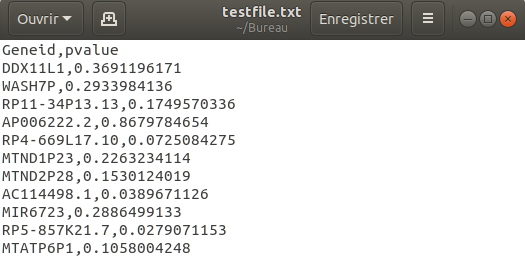
Columns :
column with gene name : 1
column with p-value : 2
Output
The most significant modules are displayed on a result page, however, all the modules found can be downloaded as an Excel document composed of two sheets. The first sheet, named "list of modules" contains the list of all modules found. The results are formated in 4 columns containing the module number, the list of genes in the module, the s score of the module and the associated p-value. The second sheet, named "genes to modules", is composed of two columns: the first one contains the name of a gene and the second one, the module to which it belongs.
Citation
Pasquier, C., Guerlais, V., Pallez, D., Rapetti-Mauss, R., & Soriani, O. (2021). Identification of active modules in interaction networks using node2vec network embedding. BioRxiv, 2021-09